
| Title | Stata 7: Performing multiple comparison tests following two-way ANOVA | |
| Author | Kenneth Higbee, StataCorp |
I am performing two-way ANOVA and wish to proceed with a multiple comparison test. Does Stata support any multiple comparison tests following two-way ANOVA?
Currently there is no official Stata command to perform multiple comparison tests for two-way (and higher order) ANOVA. It is, however, easy to construct the tests in Stata following a few simple steps:
You can either do these steps interactively in Stata, or you can write a do-file or ado-file to help automate the process.
Step (3), the correction for the multiple testing, is easy. The hardest part is step (2), performing the single degree-of-freedom test. Sometimes it can be difficult to figure out which linear combination of coefficients you are going to test. Take a look at the section titled "Testing coefficients" in [R] anova (page 59 -- Version 7 manual) and the FAQ “How can I form various tests comparing the different levels of a categorical variable after anova or regress?” for guidance on obtaining single degree-of-freedom tests.
The simplest case is when each of your single degree-of-freedom tests use the residual error from the ANOVA model as the denominator in the F tests. When your single degree-of-freedom tests do not use residual error as the denominator, step (2) requires more than one test command. See the FAQ “How can you specify a term other than residual error as the denominator in a single degree-of-freedom F test after ANOVA” for guidance.
Here is a made-up dataset:
. list
a b y
1. 1 1 4
2. 1 1 19
3. 1 1 17
4. 1 2 13
5. 1 2 9
6. 1 2 8
7. 2 1 34
8. 2 1 8
9. 2 1 11
10. 2 2 30
11. 2 2 38
12. 2 2 39
13. 3 1 39
14. 3 1 14
15. 3 1 10
16. 3 2 52
17. 3 2 51
18. 3 2 43
19. 4 1 36
20. 4 1 32
21. 4 1 50
22. 4 2 45
23. 4 2 35
24. 4 2 29
and here is the corresponding two-way ANOVA:
. anova y a b a*b
Number of obs = 24 R-squared = 0.7401
Root MSE = 9.52628 Adj R-squared = 0.6264
| Source | Partial SS df MS F Prob > F | |
| Model | 4134.50 7 590.642857 6.51 0.0010 | |
| a | 2470.16667 3 823.388889 9.07 0.0010 | |
| b | 580.166667 1 580.166667 6.39 0.0224 | |
| a*b | 1084.16667 3 361.388889 3.98 0.0270 | |
| Residual | 1452.00 16 90.75 | |
| Total | 5586.50 23 242.891304 |
The factors a and b and their interaction are significant.
Pretend that when we set up this experiment, we had decided that we wished to examine four particular individual degree-of-freedom tests on factor a. The four tests of interest for factor a are
1 and 2 versus 3 and 4
1 versus 2
3 versus 4
1, 2, and 3 versus 4
While not related to the topic of single degree-of-freedom and multiple comparison tests, looking at an interaction plot can be helpful in understanding the significant interaction from this ANOVA. One way to obtain an interaction plot is to use the graph command after the adjust command with the replace option. (preserve and restore make it easy to go to the cell mean data and then back to the original.)
. preserve . adjust , by(a b) replace
| Dependent variable: y Command: anova | ||
| b | ||
| a | 1 2 | |
| 1 | 13.3333 10 | |
| 2 | 17.6667 35.6667 | |
| 3 | 21 48.6667 | |
| 4 | 39.3333 36.3333 | |
After the adjust command with the replace option, the data now look like
. list
a b y xb
1. 1 1 4 13.33333
2. 1 2 13 10
3. 2 1 34 17.66667
4. 2 2 30 35.66667
5. 3 1 39 21
6. 3 2 52 48.66667
7. 4 1 36 39.33333
8. 4 2 45 36.33333
One way to graph the interaction plot from this data is
. sort b a . graph xb a, c(L) s([b]) xlab ylab
but I prefer
. sort b a . gen b1 = xb if b==1 . gen b2 = xb if b==2 . graph b1 b2 a, c(ll) xlab ylab psize(200)
which produces the following Stata graph:
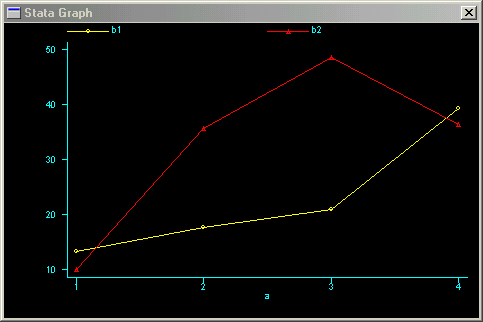
It appears that when factor a is equal to 2 or 3, the factor b effect is relatively different compared to when factor a is equal to 1 or 4.
The restore command brings us back to the original data.
. restore
Now, back to our original problem of producing multiple comparison tests. Remember that the four tests of interest for factor a are
1 and 2 versus 3 and 4
1 versus 2
3 versus 4
1, 2, and 3 versus 4
I find that using a cell means model is the easiest method for obtaining single degree-of-freedom tests. (See the FAQ How can I form various tests comparing the different levels of a categorical variable after ANOVA or regress? for details.)
The group() function of egen produces a variable indicating the cells of our design. The table command then shows the relationship between this new variable and the original a and b variables for our example.
. egen z = group(a b) . table a b, c(mean z)
| b | ||
| a | 1 2 | |
| 1 | 1 2 | |
| 2 | 3 4 | |
| 3 | 5 6 | |
| 4 | 7 8 | |
Now, I run anova using the new variable, and I make sure to specify the noconstant option.
. anova y z, noconst
Number of obs = 24 R-squared = 0.9397
Root MSE = 9.52628 Adj R-squared = 0.9095
| Source | Partial SS df MS F Prob > F | |
| Model | 22616.00 8 2827.00 31.15 0.0000 | |
| z | 22616.00 8 2827.00 31.15 0.0000 | |
| Residual | 1452.00 16 90.75 | |
| Total | 24068.00 24 1002.83333 |
(Also of interest: I could now run adjust, by(z) and see that the numbers agree with our original results.)
If I desire a Bonferroni correction on the four single degree-of-freedom tests, I simply adjust the p-values for the tests by multiplying by 4. Here is the result for comparing levels 1 and 2 against levels 3 and 4 for factor a. (Remember how the z variable relates to the original factors a and b? Level 1 of a corresponds to levels 1 and 2 of z; level 2 of a corresponds to levels 3 and 4 of z; level 3 of a corresponds to levels 5 and 6 of z; and level 4 of a corresponds to levels 7 and 8 of z.)
. test _b[z[1]]+_b[z[2]]+_b[z[3]]+_b[z[4]] = _b[z[5]]+_b[z[6]]+_b[z[7]]+_b[z[8]]
( 1) z[1] + z[2] + z[3] + z[4] - z[5] - z[6] - z[7] - z[8] = 0.0
F( 1, 16) = 19.48
Prob > F = 0.0004
. return list
scalars:
r(drop) = 0
r(df_r) = 16
r(F) = 19.48393021120293
r(df) = 1
r(p) = .0004343772345068
. di "corrected p-value = " min(1,r(p)*4)
corrected p-value = .00173751
You can see what the test command returns in r() with the command return list. We can take the uncorrected p-value of 0.00043 and produce the Bonferonni corrected p-value of .00174 by multiplying r(p) (the uncorrected p-value) by 4 (the number of tests to be performed). The test for level 1 compared to level 2 of factor a is obtained next.
. test _b[z[1]]+_b[z[2]] = _b[z[3]]+_b[z[4]]
( 1) z[1] + z[2] - z[3] - z[4] = 0.0
F( 1, 16) = 7.44
Prob > F = 0.0149
. di "corrected p-value = " min(1,r(p)*4)
corrected p-value = .05965724
Similarly, the test for level 3 compared to level 4 of factor a is obtained with
. test _b[z[5]]+_b[z[6]] = _b[z[7]]+_b[z[8]]
( 1) z[5] + z[6] - z[7] - z[8] = 0.0
F( 1, 16) = 0.30
Prob > F = 0.5930
. di "corrected p-value = " min(1,r(p)*4)
corrected p-value = 1
Notice that in this case the corrected p-value is reported as 1 since 0.5930*4 is greater than 1.
Finally, the fourth test comparing levels 1, 2, and 3 against level 4 for factor a can be obtained with the following:
. test (_b[z[1]]+_b[z[2]]+_b[z[3]]+_b[z[4]]+_b[z[5]]+_b[z[6]])/3=_b[z[7]]+_b[z[8]]
( 1) .3333333 z[1] + .3333333 z[2] + .3333333 z[3] + .3333333 z[4] +
> .3333333 z[5] + .3333333 z[6] - z[7] - z[8] = 0.0
F( 1, 16) = 8.96
Prob > F = 0.0086
. di "corrected p-value = " min(1,r(p)*4)
corrected p-value = .03435797
Other multiple comparison tests could have been performed. One popular set of multiple comparison tests for a factor with four levels are the six tests of 1 versus 2, 1 versus 3, 1 versus 4, 2 versus 3, 2 versus 4, and 3 versus 4. The same approach will work for this situation and others.