In the spotlight: Tables of estimation results in Stata 17
Making tables of estimation results is now easier than ever before with the new etable command in Stata 17. You can create tables for single or multiple models, include marginal predictions, customize your tables using the collect commands, and export your tables to most document formats. There are far too many features to demonstrate in one Stata News article. But I want to show you some basics to get you started, and you can learn more by reading the Stata manual.
Tables for one regression model
Let's begin by typing webuse nhanes2l to open a dataset based on the second National Health and Nutrition Examination Survey data (NHANES II).
. webuse nhanes2l (Second National Health and Nutrition Examination Survey)
We will use the variables diabetes, agegrp, bmi, and highbp.
. describe diabetes agegrp bmi highbp
| Variable Storage Display Value | ||
| name type format label Variable label | ||
| diabetes byte %12.0g diabetes Diabetes status | ||
| agegrp byte %8.0g agegrp Age group | ||
| bmi float %9.0g Body mass index (BMI) | ||
| highbp byte %8.0g * High blood pressure | ||
We start by fitting a logistic regression model for the binary outcome diabetes, including the predictors agegrp, bmi, and highbp. The i. prefix is factor-variable notation that tells Stata agegrp and highbp are categorical variables.
. logistic diabetes i.agegrp bmi i.highbp
Logistic regression Number of obs = 10,349
LR chi2(7) = 416.01
Prob > chi2 = 0.0000
Log likelihood = -1791.7561 Pseudo R2 = 0.1040
| diabetes | Odds ratio Std. err. z P>|z| [95% conf. interval] | |
| agegrp | ||
| 30–39 | 1.730448 .5895795 1.61 0.108 .8874554 3.374199 | |
| 40–49 | 4.259599 1.297735 4.76 0.000 2.344448 7.739213 | |
| 50–59 | 6.888277 1.993273 6.67 0.000 3.906582 12.14575 | |
| 60–69 | 10.88779 2.952805 8.80 0.000 6.398693 18.52629 | |
| 70+ | 15.25109 4.308098 9.65 0.000 8.767088 26.53057 | |
| bmi | 1.07177 .0091357 8.13 0.000 1.054014 1.089826 | |
| 1.highbp | 1.251171 .1290527 2.17 0.030 1.022161 1.531491 | |
| _cons | .0011192 .0003767 -20.19 0.000 .0005787 .0021646 | |
Now we can type etable to create a basic table of the estimation results. By default, etable displays the estimated odds ratios, their standard errors in parentheses below the odds ratios, and the number of observations.
. etable
| diabetes |
| Age group 30–39 1.730 (0.590) 40–49 4.260 (1.298) 50–59 6.888 (1.993) 60–69 10.888 (2.953) 70+ 15.251 (4.308) Body mass index (BMI) 1.072 (0.009) High blood pressure 1 1.251 (0.129) Intercept 0.001 (0.000) Number of observations 10349 |
We can use the keep() option to specify which variables to include in the table. The example below retains only the predictor variables and omits the intercept. The replay option replays the previous etable results, including any previously specified options.
. etable, replay keep(agegrp bmi highbp)
| diabetes |
| Age group 30–39 1.730 (0.590) 40–49 4.260 (1.298) 50–59 6.888 (1.993) 60–69 10.888 (2.953) 70+ 15.251 (4.308) Body mass index (BMI) 1.072 (0.009) High blood pressure 1 1.251 (0.129) Number of observations 10349 |
We can use the cstat() option to specify which statistics we wish to display in our table. The example below uses cstat(_r_b) to display the reported coefficients, which are the odds ratios for logistic regression models. The command also includes cstat(_r_ci), which displays the 95% confidence interval for the estimated odds ratios.
. etable, replay cstat(_r_b) cstat(_r_ci)
| diabetes |
| Age group 30–39 1.730 [0.887 3.374] 40–49 4.260 [2.344 7.739] 50–59 6.888 [3.907 12.146] 60–69 10.888 [6.399 18.526] 70+ 15.251 [8.767 26.531] Body mass index (BMI) 1.072 [1.054 1.090] High blood pressure 1 1.251 [1.022 1.531] Number of observations 10349 |
The table below displays the option names and descriptions of the statistics that may be used with cstat().
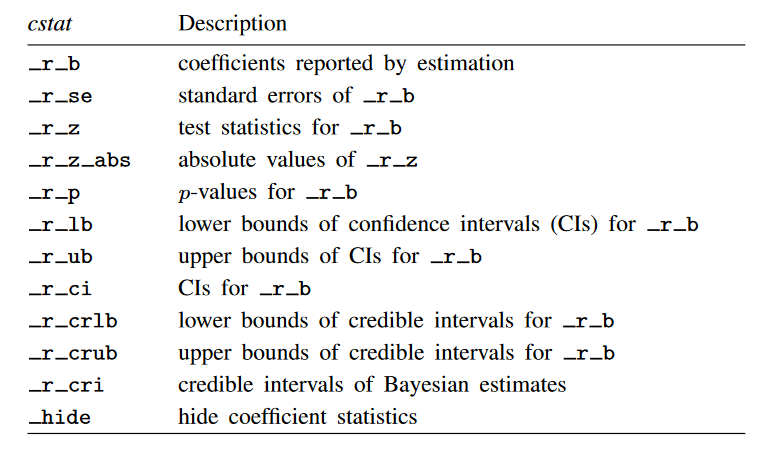

We can use the nformat() option within cstat() to specify the display format of the statistic. The example below displays the odds ratios and confidence intervals with two digits to the right of the decimal. The cidelimiter(,) option places a comma between the lower and upper bounds of the confidence interval.
. etable, replay cstat(_r_b, nformat(%4.2f))
cstat(_r_ci, cidelimiter(,) nformat(%6.2f))
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 [2.34, 7.74] 50–59 6.89 [3.91, 12.15] 60–69 10.89 [6.40, 18.53] 70+ 15.25 [8.77, 26.53] Body mass index (BMI) 1.07 [1.05, 1.09] High blood pressure 1 1.25 [1.02, 1.53] Number of observations 10349 |
We can use the showstars option to place stars next to the odds ratios with p-values below 0.05 and 0.01. We can use the showstarsnote option to place a note at the bottom of the table to remind us that one star indicates a p-value less than 0.05 and two stars indicate a p-value less than 0.01.
. etable, replay showstars showstarsnote
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 ** [2.34, 7.74] 50–59 6.89 ** [3.91, 12.15] 60–69 10.89 ** [6.40, 18.53] 70+ 15.25 ** [8.77, 26.53] Body mass index (BMI) 1.07 ** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 |
We can use the stars() option to create custom star definitions and use attach() to place the stars next to a particular statistic.
. etable, replay > showstars showstarsnote > stars(.05 "*" .01 "**" .001 "***", attach(_r_b))
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 |
We can use the mstat() option to add model-level statistics to our table. The example below uses mstat(N) to include the sample size, mstat(aic) to add Akaike's information criterion (AIC), and mstat(bic) to add Schwarz's Bayesian information criterion (BIC) to the table.
. etable, replay mstat(N) mstat(aic) mstat(bic)
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 AIC 3599.51 BIC 3657.47 |
The table below displays the identifiers and descriptions for the statistics (results) that may be used with mstat().
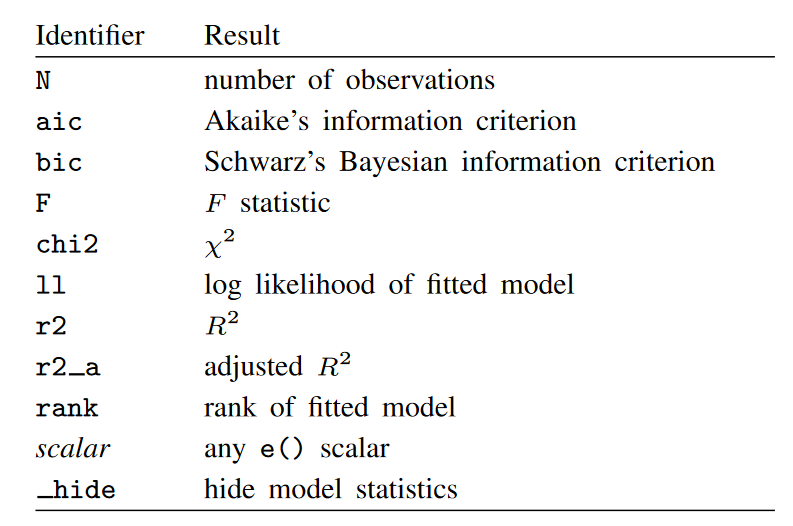

Some model-level statistics are appropriate only after a particular model. Those statistics are stored as scalars in the estimation results. You can view a list of these results by typing ereturn list immediately after you run an estimation command. And you can include those statistics by using mstat().
The scalar e(r2_p) was left in memory by the logistic command, and we use mstat(r2_p) to include it in the example below. We also use nformat() to customize the display format and label() to create a custom label for the pseudo r-squared.
. etable, replay mstat(r2_p, nformat(%5.4f) label("Pseudo R2"))
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Pseudo R2 0.1040 |
We can add titles to our tables with the title() option, and we can customize the font for the title with the titlestyles() option. Style changes affect only the title when the table is exported and will not affect the Stata output.
. etable, replay
> title("Table 5: Logistic Regression Model For Diabetes")
> titlestyles(font(Calibri, size(14) bold))
Table 5: Logistic Regression Model For Diabetes
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 AIC 3599.51 BIC 3657.47 Pseudo R2 0.1040 |
We can also use the notes option to add notes to our table, and we can customize the font with the notestyles() option. Style changes affect only the notes when the table is exported and will not affect the Stata output.
. etable, replay
> note("Data Source: NHANES, 1981")
> notestyles(font(Calibri, size(10) italic))
Table 5: Logistic Regression Model For Diabetes
| diabetes |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 AIC 3599.51 BIC 3657.47 Pseudo R2 0.1040 |
We can use the column() option to customize the column labels. The default is the name of the dependent variable, which can be specified as column(depvar). We can also use column(dvlabel) to label the column with the dependent variable label, column(command) to use the command, column(title) to use the command title stored in e(title), column(estimates) to use the names specified with estimates store, and column(index) to use sequential integers.
. etable, replay column(dvlabel)
Table 5: Logistic Regression Model For Diabetes
| Diabetes status |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 AIC 3599.51 BIC 3657.47 Pseudo R2 0.1040 |
When we are finished, we can export our table to one of many file formats by using the export() option. The example below exports our table to a Microsoft Word document named Diabetes.docx.
. etable, replay
> export(Diabetes.docx, replace)
Table 5: Logistic Regression Model For Diabetes
| Diabetes status |
| Age group 30–39 1.73 [0.89, 3.37] 40–49 4.26 *** [2.34, 7.74] 50–59 6.89 *** [3.91, 12.15] 60–69 10.89 *** [6.40, 18.53] 70+ 15.25 *** [8.77, 26.53] Body mass index (BMI) 1.07 *** [1.05, 1.09] High blood pressure 1 1.25 * [1.02, 1.53] Number of observations 10349 AIC 3599.51 BIC 3657.47 Pseudo R2 0.1040 |
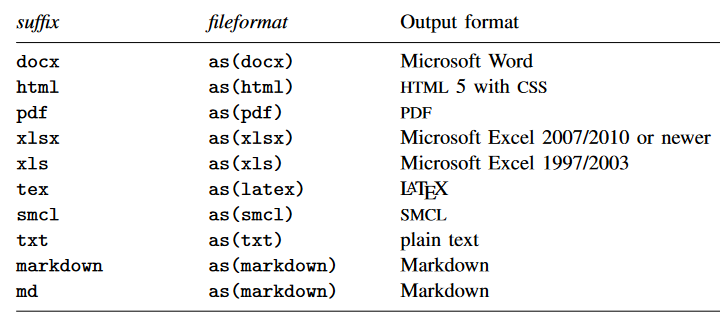
You can save your tables in any of the formats shown in the table below using the file suffixes listed.
Tables for more than one regression model
Now that we understand how to use etable for one regression model, let's see how to use etable to create tables for more than one regression model. There are several ways you can do this, and I am going to show you one way using estimates store.
We can store the results of a regression model in memory by typing estimates store name immediately after running the regression command. We can then refer to the stored results in other commands by using name. In the example below, we fit four logistic regression models and store the results of each model using estimates store. We precede each logistic command with quietly to suppress the output.
. quietly logistic diabetes i.agegrp bmi i.highbp . estimates store full . . quietly logistic diabetes i.agegrp . estimates store age . . quietly logistic diabetes bmi . estimates store bmi . . quietly logistic diabetes i.highbp . estimates store highbp
We can then include the names of our results in the estimates() option to choose the models to include in our table.
. etable, estimates(full age bmi highbp) > keep(agegrp bmi highbp)
| diabetes diabetes diabetes diabetes |
| Age group 30–39 1.730 2.017 (0.590) (0.685) 40–49 4.260 5.251 (1.298) (1.590) 50–59 6.888 9.076 (1.993) (2.596) 60–69 10.888 13.948 (2.953) (3.735) 70+ 15.251 19.494 (4.308) (5.418) Body mass index (BMI) 1.072 1.089 (0.009) (0.008) High blood pressure 1 1.251 2.577 (0.129) (0.247) Number of observations 10349 10349 10349 10349 |
In the example below, we use the column(index) option to number each of the columns and note() to add a note to list the covariates in each model.
. etable, replay column(index) > note(Model 1: agegrp bmi highbp) > note(Model 2: agegrp) > note(Model 3: bmi) > note(Model 4: highbp) > notestyles(font(Calibri, size(10) italic))
| 1 2 3 4 |
| Age group 30–39 1.730 2.017 (0.590) (0.685) 40–49 4.260 5.251 (1.298) (1.590) 50–59 6.888 9.076 (1.993) (2.596) 60–69 10.888 13.948 (2.953) (3.735) 70+ 15.251 19.494 (4.308) (5.418) Body mass index (BMI) 1.072 1.089 (0.009) (0.008) High blood pressure 1 1.251 2.577 (0.129) (0.247) Number of observations 10349 10349 10349 10349 |
Let's use all of these options together to create a final table and export the results to a Microsoft Word document named Diabetes.docx.
. etable, estimates(full age bmi highbp)
> keep(agegrp bmi highbp)
> column(index)
> stat(_r_b, nformat(%4.2f))
> cstat(_r_ci, cidelimiter(,) nformat(%6.2f))
> showstars showstarsnote
> stars(.05 "*" .01 "**" .001 "***", attach(_r_b))
> mstat(N, nformat(%8.0fc) label("Observations"))
> mstat(aic, nformat(%5.0f))
> mstat(bic, nformat(%5.0f))
> mstat(r2_p, nformat(%5.4f) label("Pseudo R2"))
> title(Table 5: Logistic Regression Model For Diabetes)
> titlestyles(font(Calibri, size(14) bold))
> note(Data Source: NHANES, 1981)
> note(Model 1: agegrp bmi highbp)
> note(Model 2: agegrp)
> note(Model 3: bmi)
> note(Model 4: highbp)
> notestyles(font(Calibri, size(10) italic))
> export(Diabetes.docx, replace)
Table 5: Logistic Regression Model For Diabetes
| 1 2 3 4 |
| Age group 30–39 1.73 2.02 * [0.89, 3.37] [1.04, 3.92] 40–49 4.26 *** 5.25 *** [2.34, 7.74] [2.90, 9.51] 50–59 6.89 *** 9.08 *** [3.91, 12.15] [5.18, 15.90] 60–69 10.89 *** 13.95 *** [6.40, 18.53] [8.25, 23.57] 70+ 15.25 *** 19.49 *** [8.77, 26.53] [11.31, 33.61] Body mass index (BMI) 1.07 *** 1.09 *** [1.05, 1.09] [1.07, 1.10] High blood pressure 1 1.25 * 2.58 *** [1.02, 1.53] [2.14, 3.11] Observations 10,349 10,349 10,349 10,349 AIC 3600 3675 3892 3900 BIC 3657 3718 3906 3915 Pseudo R2 0.1040 0.0841 0.0279 0.0258 |
Customizing etables with collect
I'm 99% happy with our final table, but I don't like the "1" label for "High blood pressure". We could use label define and label values to add a custom label to "1". But we can also customize the label without changing the original data. etable uses collect to construct tables, and we can view the underlying table structure by typing collect layout.
. collect layout
Collection: ETable
Rows: coleq#colname[agegrp bmi highbp]#result[_r_b _r_ci] result[N aic bic pseudo_r2]
Columns: cmdset#stars
Table 1: 20 x 8
(Table output omitted)
This means that we can also use collect to customize tables built with etable. In the example below, we use collect label levels to change the column name from "High blood pressure" to "Hypertension". We also label the levels of the dimension highbp.
. collect label levels colname highbp "Hypertension", modify
. collect label levels highbp 0 "No" 1 "Yes"
. collect preview
Table 5: Logistic Regression Model For Diabetes
| 1 2 3 4 |
| Age group 30–39 1.73 2.02 * [0.89, 3.37] [1.04, 3.92] 40–49 4.26 *** 5.25 *** [2.34, 7.74] [2.90, 9.51] 50–59 6.89 *** 9.08 *** [3.91, 12.15] [5.18, 15.90] 60–69 10.89 *** 13.95 *** [6.40, 18.53] [8.25, 23.57] 70+ 15.25 *** 19.49 *** [8.77, 26.53] [11.31, 33.61] Body mass index (BMI) 1.07 *** 1.09 *** [1.05, 1.09] [1.07, 1.10] Hypertension Yes 1.25 * 2.58 *** [1.02, 1.53] [2.14, 3.11] Observations 10,349 10,349 10,349 10,349 AIC 3600 3675 3892 3900 BIC 3657 3718 3906 3915 Pseudo R2 0.1040 0.0841 0.0279 0.0258 |
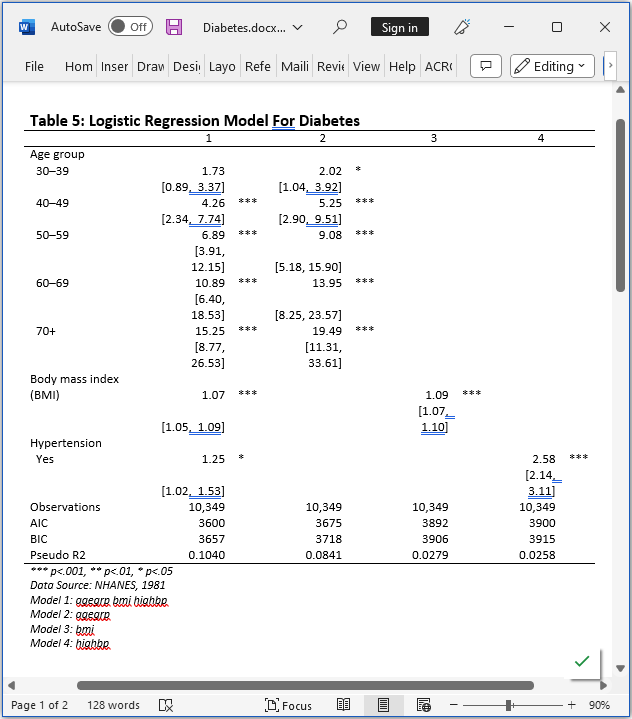
Now we can export the final table to a Microsoft Word document named Diabetes.docx by using collect style putdocx and collect export.
. collect style putdocx, layout(autofitcontents) . collect export Diabetes.docx, as(docx) replace (collection ETable exported to file Diabetes.docx)
Tables of marginal predicted probabilities using etable
You can also use etable to create tables of marginal predictions, in this case, expected probabilities of having diabetes for each group. Simply fit your model, use margins to estimate the marginal predictions, and add the margins option to etable. Here's a quick example to get you started.
. quietly logistic diabetes i.agegrp bmi i.highbp . quietly margins agegrp highbp . etable, margins
| diabetes |
| Age group 20–29 0.008 (0.002) 30–39 0.013 (0.003) 40–49 0.032 (0.005) 50–59 0.051 (0.006) 60–69 0.078 (0.005) 70+ 0.105 (0.010) High blood pressure 0 0.043 (0.003) 1 0.052 (0.003) Number of observations 10349 |
You can use the same options to customize your tables as before. You can even combine the results from your model with the marginal predictions. The example below shows how to include the odds ratios and the marginal predicted probabilities in the same table.
. quietly logistic diabetes i.agegrp bmi i.highbp . quietly etable . quietly margins agegrp highbp . etable, append margins keep(agegrp highbp) < column(index) < cstat(_r_b, nformat(%4.2f)) < cstat(_r_ci, nformat(%5.2f) cidelimiter(,)) < notes(Column 1: Odds ratios [95% CI]) < notes(Column 2: Marginal predicted probabilities [95% CI])
| 1 2 |
| Age group 20–29 0.01 [0.00, 0.01] 30–39 1.73 0.01 [0.89, 3.37] [0.01, 0.02] 40–49 4.26 0.03 [2.34, 7.74] [0.02, 0.04] 50–59 6.89 0.05 [3.91,12.15] [0.04, 0.06] 60–69 10.89 0.08 [6.40,18.53] [0.07, 0.09] 70+ 15.25 0.10 [8.77,26.53] [0.09, 0.12] High blood pressure 0 0.04 [0.04, 0.05] 1 1.25 0.05 [1.02, 1.53] [0.05, 0.06] Number of observations 10349 10349 |
Conclusion
These are just some of the fun things you can do with the new etable command in Stata 17. We've barely scratched the surface, but I hope that I've inspired you to try it out. You can learn more about etable at stata.com and in the Stata manual.
— by Chuck Huber
Director of Statistical Outreach







